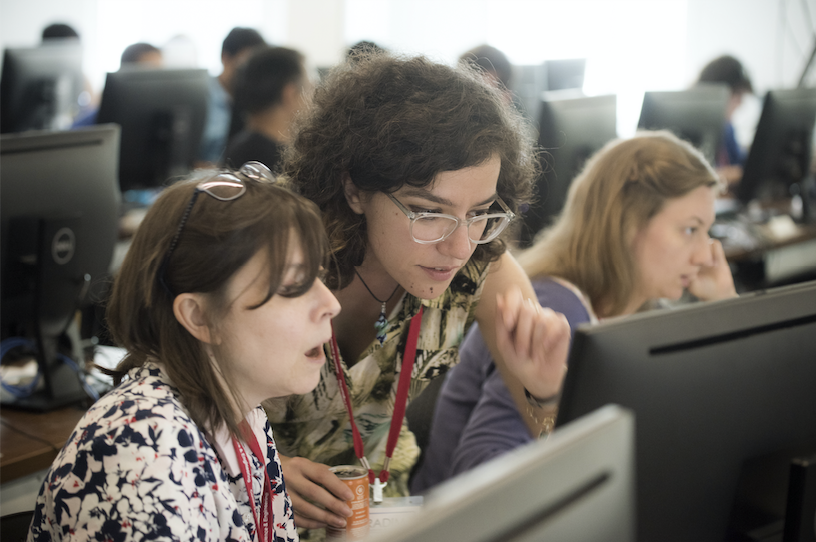
Computational resources are available as part of the theory component of PARADIM’s Materials-By-Design Toolbox. Users can find information about available software, specific hands-on tutorials, lecture videos, and a computational support forum to communicate with PARADIM scientists. The menu will navigate users to various tutorials based on standard DFT calculations using the open source Quantum Espresso package as well as many-body calculations beyond DFT using the YAMBO code. The tutorials are designing to help both expert and non-expert users. Users can also find information on how to access and run calculations at supercomputing facilities available through PARADIM.
For technical assistance with your calculations consult the PARADIM theory tutorials, or reach out to our PARADIM theory staff.
Solid State and Quantum Chemistry Codes
JDFTx is a plane-wave density-functional theory (DFT) code designed to be as easy to develop with as it is easy to use. It is distributed under the GPL license (version 3 or higher) and publications resulting from its use must cite originating publication.
Associated Publication Must cite if code is used

VASP is a computer program for atomic scale materials modeling, e.g. electronic structure calculations and quantum-mechanical molecular dynamics, from first principles. (Available through PARADIM Collaboration only)

Materials Studio is a complete modeling and simulation environment designed to allow researchers in materials science and chemistry to predict and understand the relationships of a material’s atomic and molecular structure with its properties and behavior . (Licensed to CAU and available only via collaboration with Prof. Wang)

Quantum Espresso is an Integrated suite of Open-Source computer codes for electronic-structure calculations and materials modeling at the nanoscale.

ABINIT is a program allows one to find the total energy, charge density and electronic structure of systems made of electrons and nuclei (molecules and periodic solids) within Density Functional Theory (DFT), using pseudopotentials and a plane wave or wavelet basis.

NAMD is a parallel molecular dynamics code designed for high-performance simulation of large biomolecular systems.

DFTB+ is a fast and efficient versatile quantum mechanical simulation package. It is based on the Density Functional Tight Binding (DFTB) method, containing almost all of the useful extensions which had been developed for the DFTB framework so far.

CP2K is a quantum chemistry and solid state physics software package that can perform atomistic simulations of solid state, liquid, molecular, periodic, material, crystal, and biological systems.

SIESTA is both a method and its computer program implementation to perform efficient electronic structure calculations and ab initio molecular dynamics simulations of molecules and solids.
Classical molecular dynamics code, and an acronym for Large-scale Atomic/Molecular Massively Parallel Simulator.
Visualization Codes

VESTA is a 3D visualization program for structural models, volumetric data such as electron/nuclear densities, and crystal morphologies.

Molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3-D graphics and built-in scripting.
PARADIM Data Tools and Codes

Approved PARADIM Proposals gain access to the JHU-MARCC high-performance computing center

The SciServer PARADIM Data Collective (PDC) Cloud is a collaborative research platform for large-scale data-driven science. PARADIM users can run density functional theory (DFT) computational simulations and analyze microscopy data via inbuilt Jupyter notebook recipes.

PARADIM provides access to curated data sets, public data and private user data.

Materials Automated strives to automate repetitive data analysis workflows for anyone working with materials data in materials chemistry, physics, or science
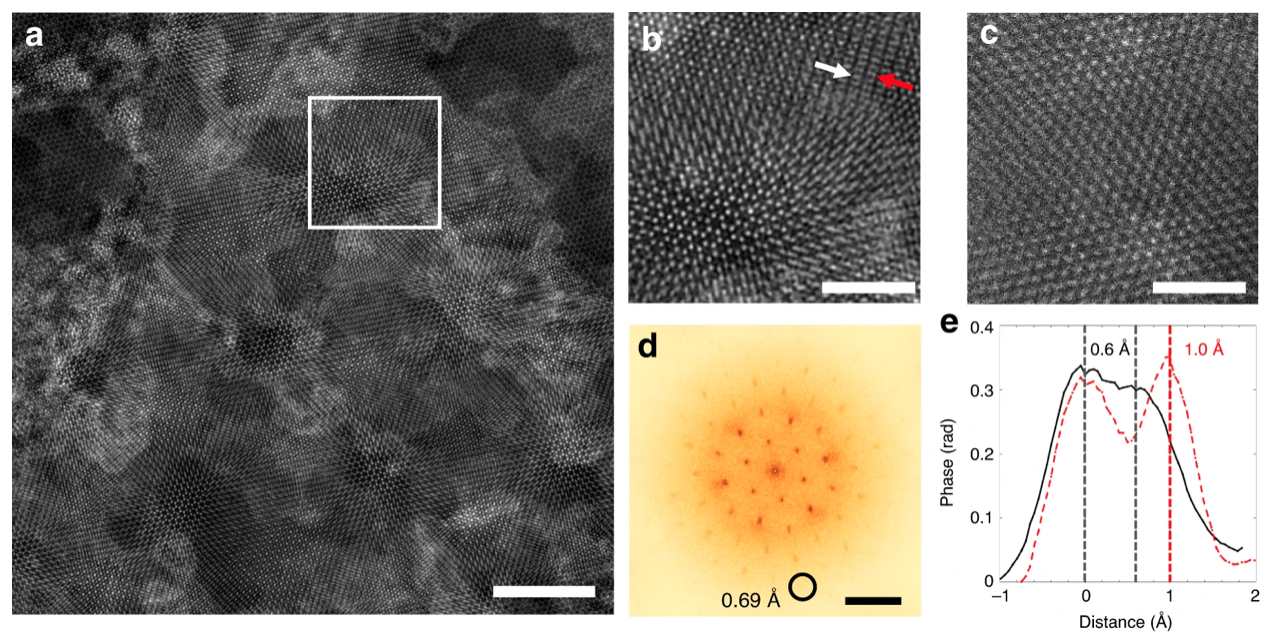
e-Ptychography is a computational imaging technique developed at Cornell University by Zhen Chen and David Muller.
Learn More
Associated Publication - Cite if code is used
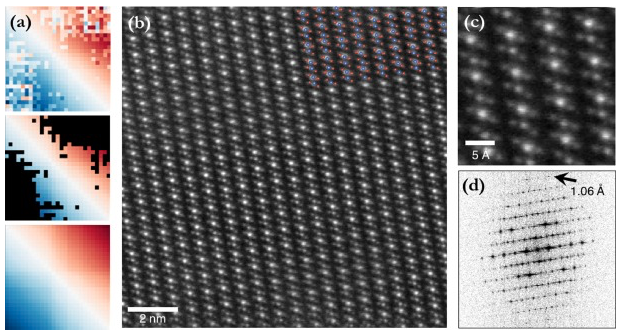

RigidRegistration is an image registration method optimized for low-SNR, cryogenic STEM data.
Learn More
Associated Publication - Must cite if code is used